mirecc:mireccanat
Table of Contents
DTI processing
- convert the raw DTI data into nifti format
- skull strip the DTI
- run eddy current correction
- run dtifit to calculate FA,eigenvalues,etc
- these steps are handled through the make_dti script
- run bedpostx on the cluster to estimate crossing fibers
- running bedpost requires copying the dtifit results to a temporary directory within the MIRECC.03/Analysis/dti_temp folder and running each slice individually through the cluster. If you do not run the slices individually they are run sequentially, which can take 1 week to finish.
- runBEDPOST_PRE.sh – sets up the bedpostx directory
- submit_CROSS.sh – submits the individual slices through runBEDPOST_CROSS.sh (this does not need to be run)
- once all 60+ slices are completed, they need to be combined into one volume and moved back to the individual subject's directory.
- navigate to the subject's folder (on Selye) and run:
bedpostx_postproc.sh dti
- copy the dti.bedpostx folder to the subject's folder on Cohen
- now dti can be normalized and dyads can be masked: post_dti_steps.sh
- navigate to /data/code and run it as bash -x post_dti_steps.sh SUBJNUM
First, FreeSurfer, Manual Comparison
- Convert the raw anatomicals into nifti, and flip them from the default orientation into LAS (FSL's preferred orientation) - script
- Run the FIRST subcortical segmentation - FIRST Methods - FSL FIRST Documentation
- Run FreeSurfer segmentation on the same T1 as FIRST - FreeSurfer Methods - FreeSurfer Documentation
- Calculate Overlap using fslstats - Overlap Methods
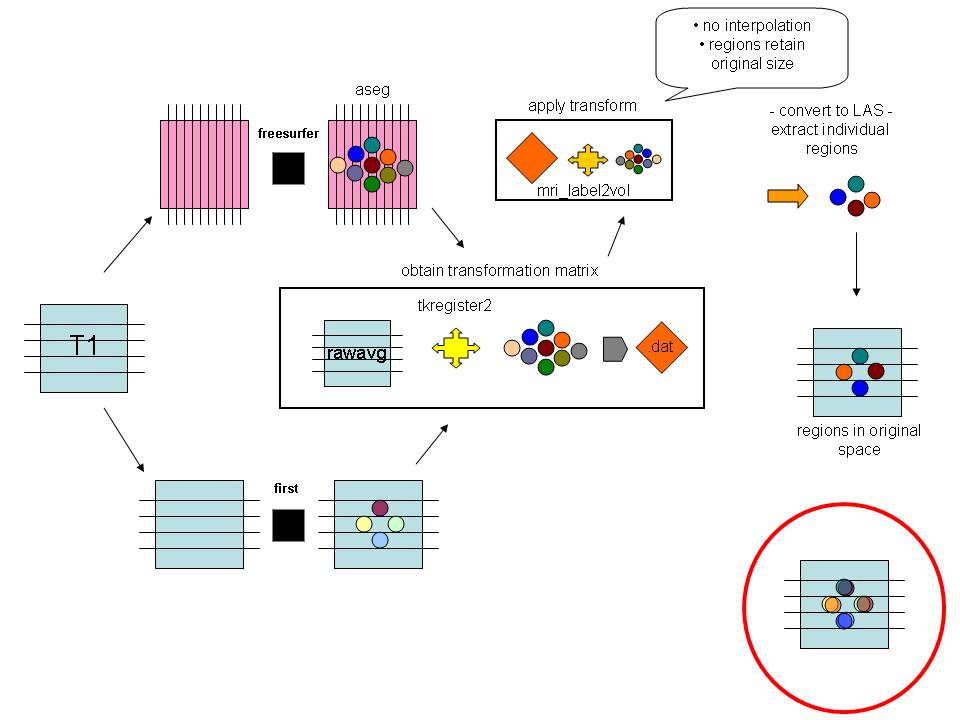
flowchart showing steps of getting first/freesurfer volumes into the same space
mirecc/mireccanat.txt · Last modified: 2024/06/21 15:44 by 127.0.0.1