User Tools
fScan – Near real-time image processing, visualization, imaging web-service
The fScan image processing program (Voyvodic, 1999, 2006; Voyvodic et al., 2009), is a single, large software program (over 100,000 lines of C code). fScan was originally developed to provide near real-time statistical image analysis, but has evolved to include a comprehensive repertoire of visualization and analysis capabilities. At Duke, fScan is used for many research and clinical fMRI applications. For patient studies, fScan’s highly automated processing capabilities have been extensively validated over the past 12 years; the accuracy of patient fMRI results are also routinely verified by direct comparison to activation map images generated using GE’s standard Brainwave software.
FScan has many innovative design features that make it well suited for patient scanning as well as for other applications including data sharing and for multi-site comparative research studies:
• Performance – An important feature of fScan is its processing speed. fScan is a single C program designed for efficiency both in terms of processing performance and memory usage. By using efficient algorithms and minimizing file I/O, the software is able to integrate a wide variety of fast processing capabilities. Fast processing enables us to routinely generate brain activation maps and head motion plots in real-time (as fast as image creation), which quickly identify many acquisition problems in time to be immediately corrected. Real-time analysis also makes scanning more efficient by allowing functional scans to be stopped as soon as a brain map reaches some analysis criterion. Although many groups have implemented different forms of on-line fMRI analysis (e.g Cox et al., 1995; Gembris et al., 2000; Posse et al., 2001; deCharms et al., 2004; Hollmann et al., 2008), the approach we have developed is particularly well suited for analysis of clinical fMRI scans because of its emphasis on quality assurance and efficient work-flow.
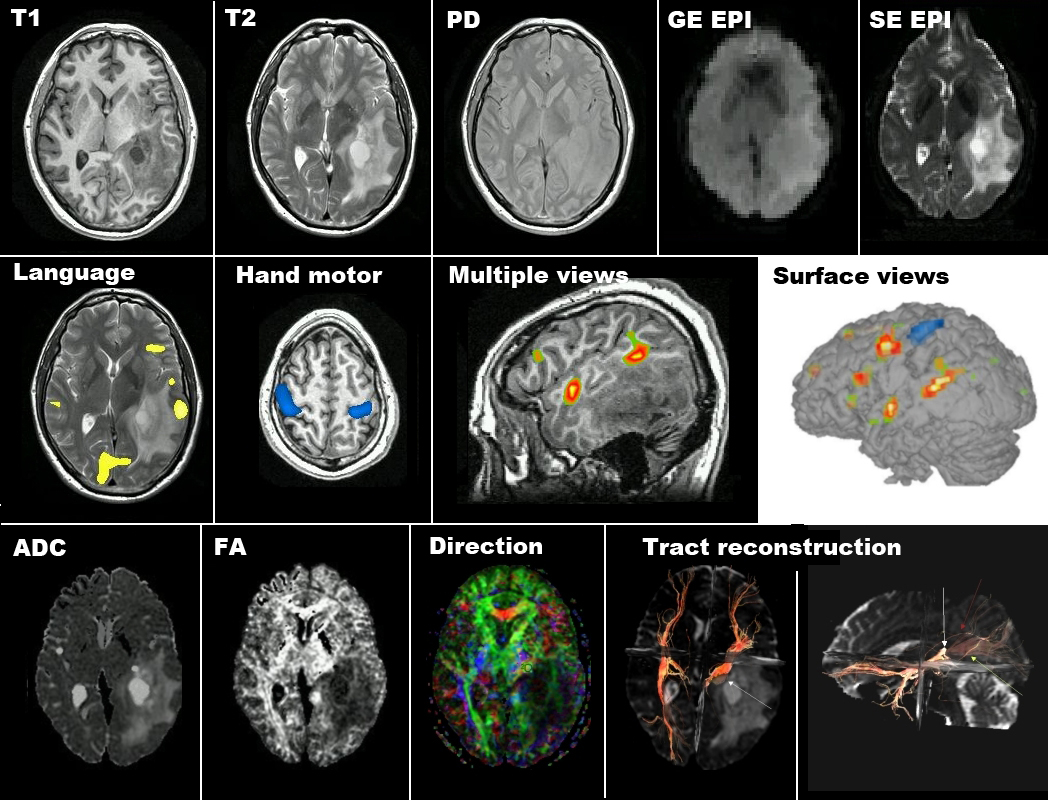
• Comprehensive analysis – Prompted by the work-flow demands of patient scans, we have adopted the strategy of incorporating all the fMRI processing tools we require into our fScan software, so that the entire analysis process can be completed on-line at the scanner console. Currently this includes real-time processing during acquisition, as well as post-processing functional scans to remove drifts, filter spikes, signal average, register images, and overlay functional maps on anatomical images. It also includes tissue segmentation and generating 3D reconstructions of the patient’s brain from anatomical images and superimposing activation maps on brain surface views. Other task-related data can also be incorporated into the analysis. Figure 1 illustrates some of fScan’s analysis and visualization features used routinely to process clinical fMRI exams.
• Workspaces – Data are organized within fScan in “io_sets” and “workspaces”. An io_set is loosely defined as a multidimensional collection of neighboring values, typically read from a single file or closely linked set of files. For example, an io_set can contain a single 2-dimensional image, or it can contain a 3-D, 4-D, 5-D or even 6-D series of images. Or it could contain a 3D surface, 3D fiber tracks, or 1-dimensional physiological time series data. The values in an io_set share a common spatial and temporal coordinate system. A workspace can contain any number of related io_sets. Each io_set within a workspace is associated with spatial transformation specifications so that within a workspace all io_sets can be aligned to a common coordinate system. Figure 2 shows an example of multiple io_sets in a single workspace. Two complete workspaces can be loaded in fScan simultaneously. When two io_sets from different workspaces are then registered to each other, any spatio-temporal point in any io_set can be mapped to the corresponding point in any other io_set. The number of io_sets that can be simultaneously loaded into fScan is typically not limited by the amount of available computer memory. This is because fScan’s memory management is dynamically controlled so that only active images are stored in memory; inactive data buffers are purged automatically when more memory is needed. Newly created io_sets may need to be written out to disk files to facilitate efficient memory usage.
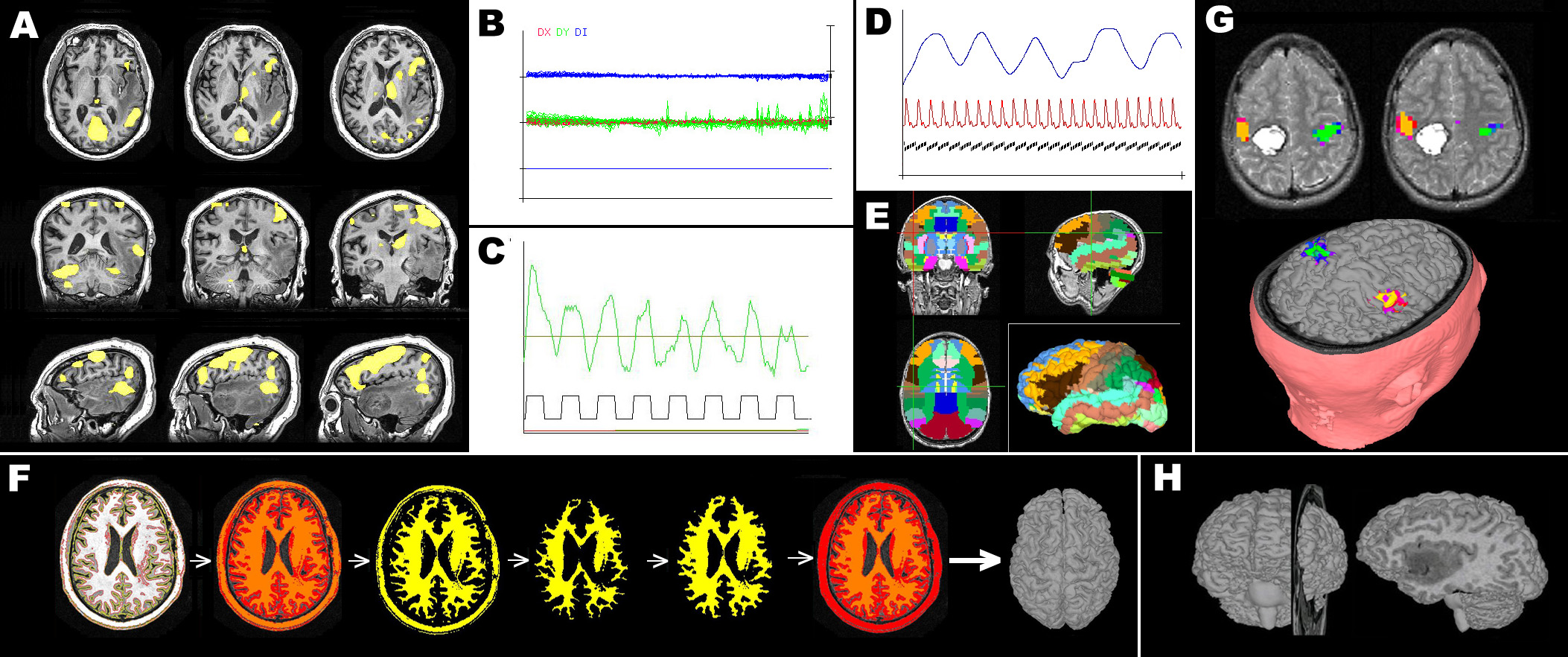
Figure 2. Examples of fScan analysis features. A) Statistical activation maps generated as t-maps and overlaid on any image (in this case T1 images resliced in 3 planes). B) Head motion (green & red) and image intensity (blue) are plotted together in stability plots. T-maps and stability plots are routinely generated in near real-time during image acquisition. C) Image intensity fluctuations are plotted and compared to task reference plots. D) Behavioral and physiological data are automatically temporally registered with image timing. Here respiration and heart beat are shown with an image timing reference (black ticks are MRI images). E) A reference atlas such as the Wake Forest PickAtlas can be registered with a patient brain to create anatomical ROIs. F) T1 images can be semi-automatically segmented and reconstructed to brain surfaces. The sequence shown took less than 30s to complete (actual fScan command sequence is shown in Figure 3). G) Activation maps can be painted onto brain surfaces and multiple reconstructions (e.g. brain plus head as shown) can be combined. H) Surface reconstructions can be combined with cutting planes and MRI images to reslice the brain in any orientation.
• Flexibility – fScan features hundreds of data processing and visualization commands that can either be entered interactively or via scripted command files and menus. Its internal command repertoire includes simple image processing primitives (e.g. add, subtract, multiply, erode, dilate), as well as more sophisticated statistical operations such as t-tests, fast Fourier transforms, and 3-D surface reconstructions. The scripting and programmable graphical user interface (GUI) features provide enormous flexibility for organizing analysis and visualization protocols tailored for different applications. GUI menus, in particular, provide a simple way to create highly automated customized operations that can be executed with a single mouse click (Figure. Flexibility is also provided by fScan’s ability to handle many different types of data, including raw images, statistical maps, behavioral timing, physiological recordings, 3D surface data, region of interest (ROI) specifications, transformation matrices, and deformation maps. All spatial and temporal data are referenced to common workspace coordinates so that locations and times can be compared across different data modalities.
• Interoperability – fScan’s processing capabilities are not limited to its own internally programmed features. It also supports in-line execution of other software via pre-packaged shell command files, run using fScan’s ‘execute’ command (see Figure 3). For example, to use FSL’s motion correction program (MCFLIRT) a user simply needs to click inside the window of the data set to be processed and then click on the “FSL MC” icon in the “stats” taskbar. A new process is started for a shell script that automatically reformats the appropriate data file to NIfTI format if necessary, runs the MCFLIRT program, and returns a command string to the waiting fScan process to automatically load the newly created data set into a new fScan window. This shell scripting option provides enormous flexibility to combine fScan’s own internal features with complementary features available in other software packages. If desired, fScan could be used simply as a GUI interface to run other programs.
• Compatibility – Another feature related to interoperability is data format compatibility. fScan can read and write a wide variety of different commonly used data file formats, including: BXH/XCEDE, NIfTI, Analyze, DICOM, JPEG, TIFF, as well as the various file formats used by the FreeSurfer software package such as COR, MGZ, surface files, and paint files. It can also read behavioral timing files in a variety of formats including FSL schedule files, XCEDE event files, and the multi-record archive files that CIGAL produces. Although the default data file format for fScan is the BXH/XCEDE generic data descriptor, the ability to read and write other formats directly facilitates direct interoperability with MRI scanners and other software.
• Portability – We have implemented native code versions of fScan for X-based operating systems (Linux, Unix, AIX, Mac OS-X) and for Windows™ operating systems. Supporting multiple platform is especially important to give remote users access to the federated imaging databases via the web, as described below.
• Simplicity – The taskbar GUI interface is designed to be intuitively easy to use, and fScan’s help features are simple and always available. Documentation is written in a web-based Wiki so that it can be easily edited, and fScan’s internal “help” command displays pages directly from the Wiki web-site so that they are always up-to-date. A local copy of the Wiki pages is also provided for documentation when fScan is used off-line.
To see how fScan's design differs from other programs, see fScan's Differences.